Biomedical Language Model
LLM-powered rare disease recognition from NIH-funded research using BioMedBERT, fine-tuned transformers, and graph analytics in Neo4j.
1. Introduction
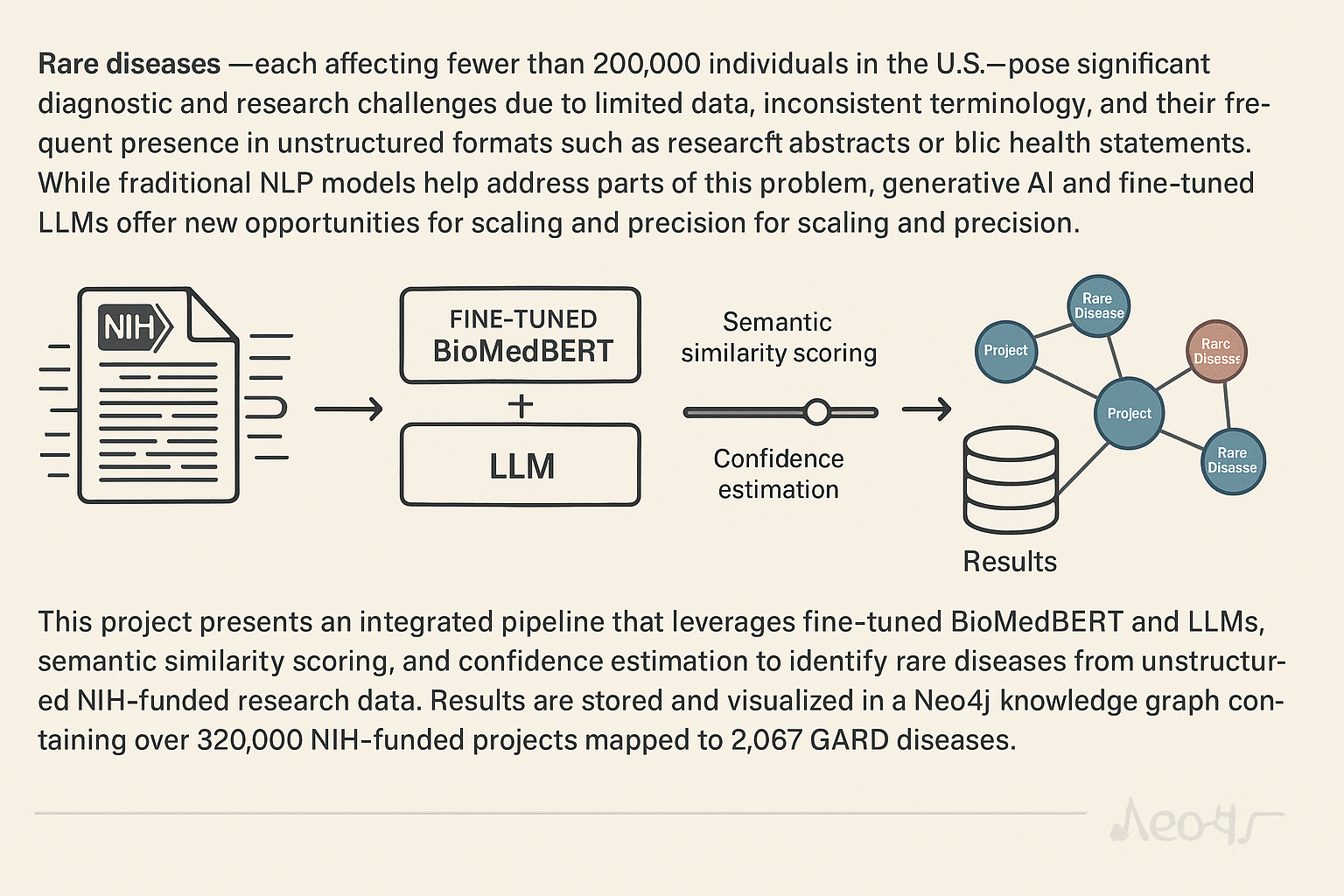
Rare diseases—each affecting fewer than 200,000 individuals in the U.S.—pose significant diagnostic and research challenges due to limited data, inconsistent terminology, and their frequent presence in unstructured formats such as research abstracts or public health statements. While traditional NLP models help address parts of this problem, generative AI and fine-tuned LLMs offer new opportunities for scaling and precision.
This project presents an integrated pipeline that leverages fine-tuned BioMedBERT and LLMs, semantic similarity scoring, and confidence estimation to identify rare diseases from unstructured NIH-funded research data. Results are stored and visualized in a Neo4j knowledge graph containing over 320,000 NIH-funded projects mapped to 2,067 GARD diseases.
2. Key Questions Addressed
- How can LLMs enhance rare disease recognition from free-text biomedical data?
- How do semantic similarity and contextual confidence scores improve precision?
- What methods allow large-scale, explainable, and queryable discovery across NIH grants?
- How can this work accelerate collaborations, drug discovery, and funding transparency?
3. The Problem
Rare disease identification in unstructured biomedical texts presents several challenges:
- Data Scarcity & Fragmentation: Most rare disease knowledge exists in unstructured, unindexed formats.
- Terminological Complexity: Rare diseases have multiple synonyms and informal references.
- Model Limitations: Pretrained general-domain LLMs struggle to capture rare disease semantics without adaptation.
- Precision vs. Recall: Simple string matching causes false positives; generative models require task-specific guidance.
4. The Importance
This work contributes a generative AI-driven framework that:
- Identifies high-value rare disease projects through language modeling and fine-grained linguistic analysis.
- Maps unstructured NIH project data into a searchable, visual knowledge graph.
- Evaluates fine-tuned LLM performance using biomedical benchmark tasks and LightEval metrics.
- Equips researchers and institutions with tools for collaboration, funding insights, and data-driven discovery.
5. The Solution
5.1. Data Sources
- NIH RePORTER API: Titles, abstracts, public health statements, and cost data from over 75,000 funded projects.
- GARD, OMIM, Orphanet: Rare disease names, synonyms, and structured definitions.
- Custom & Benchmark Datasets: Six curated datasets for training/fine-tuning BioMedBERT and LLMs.
5.2. Disease Identification & Relevance Assessment
- Exact String Matching: Enhanced with stemming, synonym expansion, and bag-of-words variants.
- Confidence Score: Weighted scoring based on mention location (title, abstract, relevance statement), sentence tense, and negation handling.
- Semantic Similarity Score: Measured using cosine similarity between contextual embeddings from a fine-tuned BioMedBERT model.
5.3. LLM Fine-Tuning and Evaluation
- LLM Training on Amazon SageMaker: Scalable model training, experimentation, and deployment using rare disease text corpora.
- Benchmarking via LightEval & MMLU: Models evaluated across:
- College Medicine
- College Biology
- Medical Genetics
- Clinical Knowledge
- High School Biology
- Anatomy
- Domain Focus: Leaderboard tasks like
mmlu:College_medicine|5|0,mmlu:Medical_genetics|5|0were emphasized to validate medical domain understanding.
5.4. Knowledge Graph Construction
- Nodes & Relationships: Over 320,000 NIH-funded projects linked to 2,067 rare diseases in Neo4j.
- Search & Query: Cypher-based querying of disease-focus relationships, funding trends, and publication impact.
- Public Access: Graph exposed at grants4rd.ncats.io.
6. Results and Outcomes
- Semantic Similarity (Relevant vs. Irrelevant): 0.81 vs. 0.50
- Confidence Score (Relevant vs. Irrelevant): 0.83 vs. 0.56
- Combined Score Model Accuracy: 0.78
- Graph Stats: 75,373 projects, 145k+ abstracts, 135k+ public health statements
- LLM Benchmarking: Strong performance across LightEval/MMLU tasks with domain-specific fine-tuning
7. Case Studies
- Funding Landscape Analysis: Mapped rare disease research trends by state and institution; strong correlation (r = 0.97) between funding and publication output.
- Hemophilia B Research: Extracted 2,301 relevant projects with detailed drug development timelines and biological targets.
- Primary Sclerosing Cholangitis (PSC): Identified under-explored grant areas and candidate repurposing opportunities.
8. Skills and Tools Used
- LLMs & NLP Models: BioMedBERT, Bio_ClinicalBERT, BlueBERT, GPT-3.5, LightEval, HuggingFace Transformers
- Model Training & Deployment: Amazon SageMaker, PyTorch, AdamW Optimizer
- Biomedical Datasets: GARD, OMIM, Orphanet, NIH RePORTER
- Evaluation & Benchmarking: MMLU domain tasks, semantic similarity, accuracy, F1-score
- Graph Analytics: Neo4j, Cypher, rdas_gfkg browser
- Visualization: Tableau, matplotlib
- Programming & Processing: Python, spaCy, scikit-learn, pandas, NumPy
9. Future Directions
- Multi-Modal Integration: Incorporate genomic, phenotypic, and clinical trial data.
- Real-Time LLMs for Research Triage: Use prompt-based agents for project summarization and disease relevance classification.
- Expanded Graph Coverage: Integrate funding data from Stanford, Harvard, Duke, and CZI.
- Advanced LLM Tuning: Explore OrphaGPT, Llama-3, and Retrieval-Augmented Generation for disease-specific knowledge extraction.
- Collaborator Recommendation Engine: Leverage graph and LLM embeddings to match investigators and institutions by focus area.