Advanced RAG & Flow Enginering
AI-powered social knowledge graph, integrating LLMs, RAG, and clustering pipelines to connect and recommend rare disease specialists.
1. Introduction
Rare diseases (RDs) affect millions globally, yet research efforts remain fragmented due to limited collaboration and access to specialists. Patients often face long diagnostic delays, and researchers may work in isolation without access to peers with complementary expertise. To address these issues, we developed a Rare Disease Research Collaborative Network (RCN) that integrates data from PubMed, NIH grants, and clinical trials into a knowledge graph using Neo4j.
By leveraging social network analysis and large language models (LLMs), we cluster researchers based on shared contributions and extract their expertise using prompt-based summarization methods. The result is a labeled, clustered RCN (lc-RCN) that supports both collaboration and specialist recommendation.
2. Key Questions Addressed
- How can social network analysis and LLMs improve researcher collaboration in rare disease research?
- What are the optimal strategies for clustering and summarizing researcher expertise?
- How can we generate fine-tuned and explainable expert recommendations based on user queries?
- What evaluation methods best capture the quality of expert extraction and summarization?
3. The Problem
Rare disease research is hampered by several issues:
- Fragmented Networks: Researchers often work in silos without awareness of nearby or relevant collaborators.
- Diagnostic Delays: Patients lack access to researchers with relevant expertise.
- Limited Labelled Data: Difficulty in labeling expert domains at scale.
- Suboptimal Collaboration Tools: Existing platforms don’t integrate modern AI-driven insights or local geographic mapping.
4. The Importance
By automating the discovery of researcher expertise and mapping collaborative relationships, RCN:
- Enhances RD research by fostering interdisciplinary connections.
- Improves patient outcomes via access to informed specialists.
- Informs funding and policy decisions through data-driven network visualization.
- Enables targeted exploration of research themes and gaps.
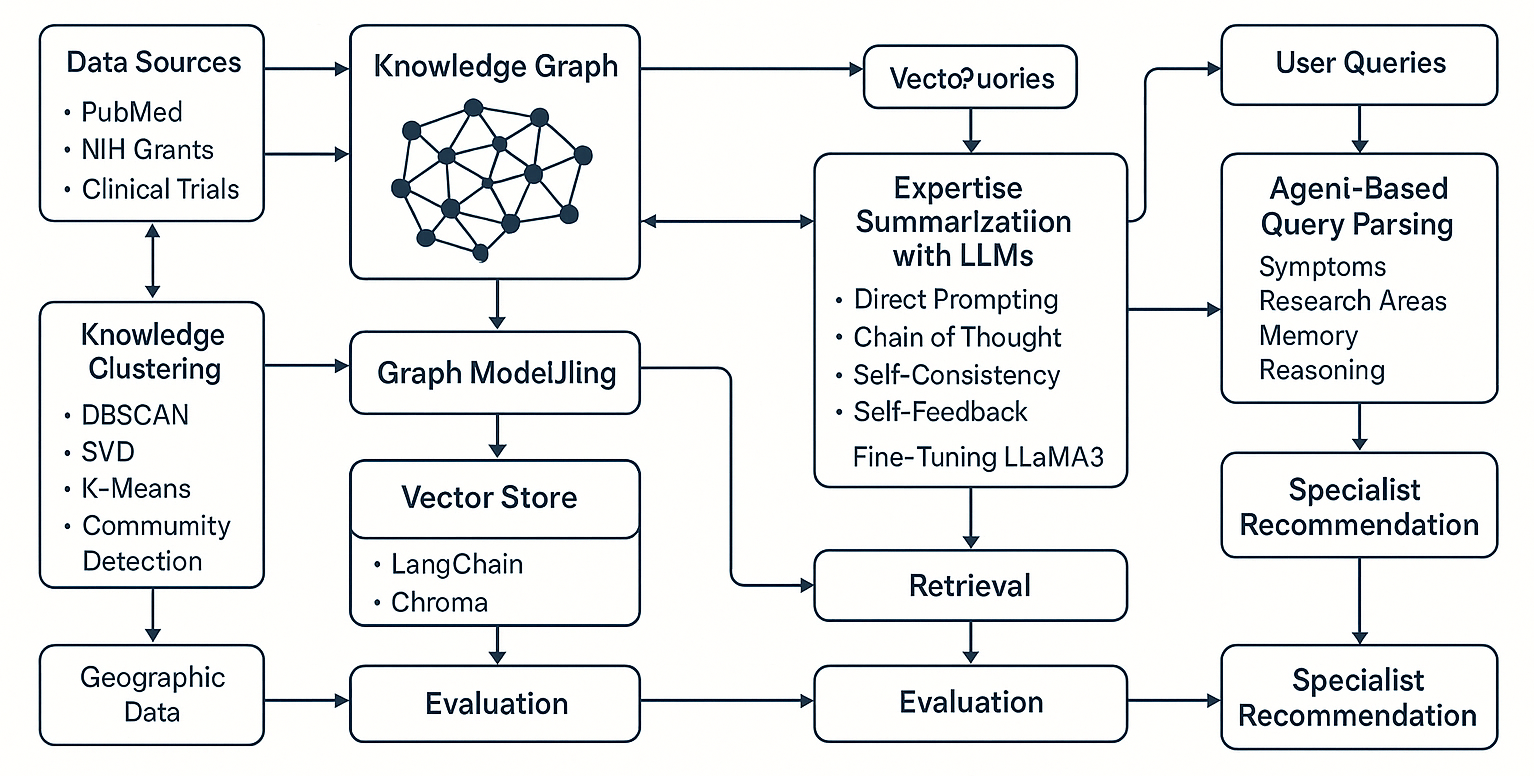
5. The Solution
5.1 Data Collection and Graph Modeling
- Collected over 6M PubMed authors, 76K grant PIs, and 100K clinical trial investigators from RDAS.
- Constructed a knowledge graph in Neo4j using nodes for researchers, projects, trials, and publications.
- Added geographic data using FIPS and coordinates to support proximity-based collaboration.
5.2 Researcher Clustering (c-RCN)
- Applied DBSCAN with SVD-based dimensionality reduction on research abstracts.
- Compared clustering against alternatives like K-Means and community detection.
5.3 Expertise Summarization with LLMs (lc-RCN)
- Tested various prompt engineering methods:
- Direct prompting
- Chain of Thought (CoT)
- Self-Consistency CoT
- Self-Feedback with iterative refinement
- Fine-tuned LLaMA 3 using LoRA for PEFT-based adaptation.
- Enhanced inference speed with Key-Value Caching.
5.4 Prompt Engineering Evaluation
- Evaluated output quality with metrics: ROUGE, BERTScore, MoverScore, BLANC, and QuestEval.
- CoT outperformed others in both semantic similarity and factual fidelity.
6. Specialist Recommendation Pipeline
6.1 Retrieval-Augmented Generation (RAG)
- Stored summarized expertise in a vector store using Langchain + Chroma.
- Queried vector store and retrieved matched researchers using Neo4j Cypher.
- Generated final recommendations through augmented prompts.
6.2 Agent-Based Query Parsing
- Built agents using langchain_core to parse open-ended patient/researcher queries.
- Extracted relevant symptoms or research areas, mapped them to expertise keywords, and retrieved specialists.
- Added memory and reasoning to handle informal input like:
“I’ve been having muscle weakness and liver dysfunction. Any idea what’s going on?”
7. Results and Impact
- Constructed a graph with 3 million researchers, 12,354 labeled expertise areas.
- Clustered RD researchers into 3,927 groups with unique expertise tags.
- Recommended specialists with high semantic similarity (>0.79 BERTScore).
- Discovered key collaboration hubs and isolated nodes.
- Enabled U.S.-based geographic mapping using affiliation FIPS codes.
8. Skills and Tools Used
Large Language Models (LLMs)
- OpenAI GPT (GPT-3.5-turbo, GPT-4o)
- Meta LLaMA-3 (PEFT + LoRA)
- Claude, Gemini, Phi, Mistral, DeepSeek
Prompt Engineering & Fine-Tuning
- SmartLLMChain, Chain-of-Thought (CoT), Self-Consistency, Self-Feedback
- Customized expert summarization with RAG pipelines
- LoRA-based fine-tuning using Unsloth and HuggingFace
Cloud Computing and Scalability
- Amazon SageMaker for scalable LLM inference, training jobs, and experiment tracking
- AWS EC2 for heavy lifting in graph analytics and visualization
- Neo4j AuraDB for managed graph database integration
Retrieval-Augmented Generation (RAG)
- Langchain + Chroma for vector search
- Cypher-based real-time specialist retrieval from Neo4j
NLP Evaluation and Metrics
- ROUGE, BERTScore, MoverScore, BLANC, QuestEval
Data Science & Preprocessing
- Pandas, Scikit-learn, SpaCy, NLTK
- Clustering via DBSCAN, dimensionality reduction via SVD
Visualization & Analysis
- NetworkX, Matplotlib for social graphs and cluster maps
- PageRank, hub/authority scoring, and geographic mapping
Deployment
- Dockerized workflow for reproducibility
- APIs for Gradio-based UIs and Langchain agents
9. Future Directions
- Incorporate PubMedBERT with STSB + GARD fine-tuning for better similarity scoring.
- Build a web UI to support patient queries and matchmaking.
- Integrate location-aware filters for proximity-based specialist recommendation.
- Explore multi-modal data integration (e.g., OMIM, genetic profiles).
- Improve name disambiguation and organizational record linkage.
10. Generative AI Capabilities
This project exemplifies the generative AI lifecycle for domain-specific data science use cases in rare disease research:
- Contextual Embedding Generation: Summarizes research background using LLMs via custom prompts
- Information Retrieval: Retrieves experts with semantically matched embeddings
- Text Generation and Summarization: Converts structured knowledge into natural language responses
- Self-Improving Feedback Loops: Uses self-feedback agents to iteratively refine keyword outputs
- Performance Evaluation: Evaluates generation outputs with multiple semantic and factual metrics
- Data Science Integration: Uses statistical, clustering, and geographic methods to analyze collaboration dynamics